In biology, histones are highly basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei and in most Archaeal phyla. They act as spools around which DNA winds to create structural units called nucleosomes.[1][2] Nucleosomes in turn are wrapped into 30-nanometer fibers that form tightly packed chromatin. Histones prevent DNA from becoming tangled and protect it from DNA damage. In addition, histones play important roles in gene regulation and DNA replication. Without histones, unwound DNA in chromosomes would be very long. For example, each human cell has about 1.8 meters of DNA if completely stretched out; however, when wound about histones, this length is reduced to about 90 micrometers (0.09 mm) of 30 nm diameter chromatin fibers.[3]
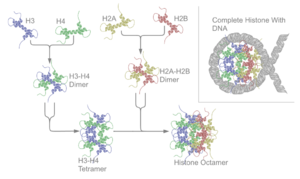
There are five families of histones which are designated H1/H5 (linker histones), H2, H3, and H4 (core histones). The nucleosome core is formed of two H2A-H2B dimers and a H3-H4 tetramer. The tight wrapping of DNA around histones is to a large degree a result of electrostatic attraction between the positively charged histones and negatively charged phosphate backbone of DNA.
Histones may be chemically modified through the action of enzymes to regulate gene transcription. The most common modification are the methylation of arginine or lysine residues or the acetylation of lysine. Methylation can affect how other protein such as transcription factors interact with the nucleosomes. Lysine acetylation eliminates a positive charge on lysine thereby weakening the electrostatic attraction between histone and DNA resulting in partial unwinding of the DNA making it more accessible for gene expression.
- ^ Youngson RM (2006). Collins Dictionary of Human Biology. Glasgow: HarperCollins. ISBN 978-0-00-722134-9.
- ^ Cox M, Nelson DR, Lehninger AL (2005). Lehninger Principles of Biochemistry. San Francisco: W.H. Freeman. ISBN 978-0-7167-4339-2.
- ^ Redon C, Pilch D, Rogakou E, Sedelnikova O, Newrock K, Bonner W (April 2002). "Histone H2A variants H2AX and H2AZ". Current Opinion in Genetics & Development. 12 (2): 162–9. doi:10.1016/S0959-437X(02)00282-4. PMID 11893489.