This article needs additional citations for verification. (January 2010) |
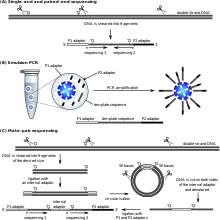

SOLiD (Sequencing by Oligonucleotide Ligation and Detection) is a next-generation DNA sequencing technology developed by Life Technologies and has been commercially available since 2006. This next generation technology generates 108 - 109 small sequence reads at one time. It uses 2 base encoding to decode the raw data generated by the sequencing platform into sequence data.
This method should not be confused with "sequencing by synthesis," a principle used by Roche-454 pyrosequencing (introduced in 2005, generating millions of 200-400bp reads in 2009), and the Solexa system (now owned by Illumina) (introduced in 2006, generating hundreds of millions of 50-100bp reads in 2009)
These methods have reduced the cost from $0.01/base in 2004 to nearly $0.0001/base in 2006 and increased the sequencing capacity from 1,000,000 bases/machine/day in 2004 to more than 5,000,000,000 bases/machine/day in 2009. Over 30 publications exist describing its use first for nucleosome positioning from Valouev et al.,[1] transcriptional profiling or strand sensitive RNA-Seq with Cloonan et al.,[2] single cell transcriptional profiling with Tang et al.[3] and ultimately human resequencing with McKernan et al.[4]
The method used by this machine (sequencing-by-ligation) has been reported to have some issue sequencing palindromic sequences.[5]
- ^ Valouev A, Ichikawa J, Tonthat T, et al. (July 2008). "A high-resolution, nucleosome position map of C. elegans reveals a lack of universal sequence-dictated positioning". Genome Research. 18 (7): 1051–63. doi:10.1101/gr.076463.108. PMC 2493394. PMID 18477713.
- ^ Cloonan N, Forrest AR, Kolle G, et al. (July 2008). "Stem cell transcriptome profiling via massive-scale mRNA sequencing". Nature Methods. 5 (7): 613–9. doi:10.1038/nmeth.1223. PMID 18516046. S2CID 19790151.
- ^ Tang F, Barbacioru C, Wang Y, et al. (May 2009). "mRNA-Seq whole-transcriptome analysis of a single cell". Nature Methods. 6 (5): 377–82. doi:10.1038/nmeth.1315. PMID 19349980. S2CID 16570747.
- ^ McKernan KJ, Peckham HE, Costa GL, et al. (September 2009). "Sequence and structural variation in a human genome uncovered by short-read, massively parallel ligation sequencing using two-base encoding". Genome Research. 19 (9): 1527–41. doi:10.1101/gr.091868.109. PMC 2752135. PMID 19546169.
- ^ Yu-Feng Huang; Sheng-Chung Chen; Yih-Shien Chiang & Tzu-Han Chen (2012). "Palindromic sequence impedes sequencing-by-ligation mechanism". BMC Systems Biology. 6 (Suppl 2): S10. doi:10.1186/1752-0509-6-S2-S10. PMC 3521181. PMID 23281822.