| Aldehyde dehydrogenase (NAD+) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
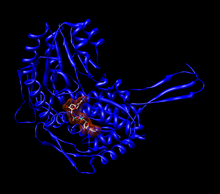 Monomer of human aldehyde dehydrogenase 2 (ALDH2) with a space-filling model of NAD+ in the active site.[1] | |||||||||
| Identifiers | |||||||||
| EC no. | 1.2.1.3 | ||||||||
| CAS no. | 9028-86-8 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Aldehyde dehydrogenases (EC 1.2.1.3) are a group of enzymes that catalyse the oxidation of aldehydes.[2] They convert aldehydes (R–C(=O)–H) to carboxylic acids (R–C(=O)–O–H). The oxygen comes from a water molecule. To date, nineteen ALDH genes have been identified within the human genome. These genes participate in a wide variety of biological processes including the detoxification of exogenously and endogenously generated aldehydes.
- ^ Cite error: The named reference
pmid12795606was invoked but never defined (see the help page). - ^ Marchitti SA, Brocker C, Stagos D, Vasiliou V (June 2008). "Non-P450 aldehyde oxidizing enzymes: the aldehyde dehydrogenase superfamily". Expert Opinion on Drug Metabolism & Toxicology. 4 (6): 697–720. doi:10.1517/17425255.4.6.697. PMC 2658643. PMID 18611112.