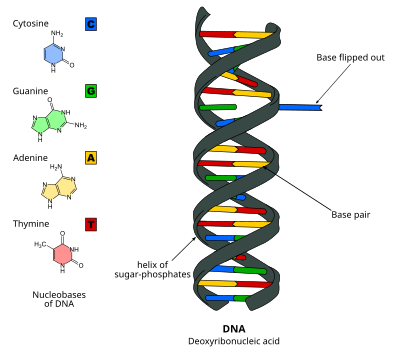
DNA base flipping, or nucleotide flipping, is a mechanism in which a single nucleotide base, or nucleobase, is rotated outside the nucleic acid double helix.[1] This occurs when a nucleic acid-processing enzyme needs access to the base to perform work on it, such as its excision for replacement with another base during DNA repair. It was first observed in 1994 using X-ray crystallography in a methyltransferase enzyme catalyzing methylation of a cytosine base in DNA. Since then, it has been shown to be used by different enzymes in many biological processes such as DNA methylation, various DNA repair mechanisms, and DNA replication. It can also occur in RNA double helices [2] or in the DNA:RNA intermediates formed during RNA transcription.
DNA base flipping occurs by breaking the hydrogen bonds between the bases and unstacking the base from its neighbors. This could occur through an active process, where an enzyme binds to the DNA and then facilitates rotation of the base, or a passive process, where the base rotates out spontaneously, and this state is recognized and bound by an enzyme. It can be detected using X-ray crystallography, NMR spectroscopy, fluorescence spectroscopy, or hybridization probes.
- ^ Roberts, RJ; Cheng, X (1998). "Base flipping". Annual Review of Biochemistry. 67 (1): 181–198. doi:10.1146/annurev.biochem.67.1.181. PMID 9759487.
- ^ Reiter, NJ; Blad, H; Abildgaard, F; Butcher, SE (2004). "Dynamics in the U6 RNA intramolecular stem-loop: a base flipping conformational change". Biochemistry. 43 (43): 13739–47. doi:10.1021/bi048815y. PMID 15504036. S2CID 25391616.