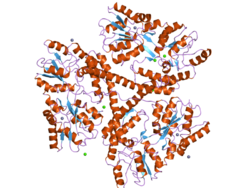
Huntingtin (Htt) is the protein coded for in humans by the HTT gene, also known as the IT15 ("interesting transcript 15") gene.[5] Mutated HTT is the cause of Huntington's disease (HD), and has been investigated for this role and also for its involvement in long-term memory storage.[6]
It is variable in its structure, as the many polymorphisms of the gene can lead to variable numbers of glutamine residues present in the protein. In its wild-type (normal) form, the polymorphic locus contains 6-35 glutamine residues. However, in individuals affected by Huntington's disease (an autosomal dominant genetic disorder), the polymorphic locus contains more than 36 glutamine residues (highest reported repeat length is about 250).[7] Its commonly used name is derived from this disease; previously, the IT15 label was commonly used.
The mass of huntingtin protein is dependent largely on the number of glutamine residues it has; the predicted mass is around 350 kDa. Normal huntingtin is generally accepted to be 3144 amino acids in size. The exact function of this protein is not known, but it plays an important role in nerve cells. Within cells, huntingtin may or may not be involved in signaling, transporting materials, binding proteins and other structures, and protecting against apoptosis, a form of programmed cell death. The huntingtin protein is required for normal development before birth.[8] It is expressed in many tissues in the body, with the highest levels of expression seen in the brain.
- ^ a b c GRCh38: Ensembl release 89: ENSG00000197386 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000029104 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ The Huntington's Disease Collaborative Research Group (Mar 1993). "A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. The Huntington's Disease Collaborative Research Group" (PDF). Cell. 72 (6): 971–83. doi:10.1016/0092-8674(93)90585-E. hdl:2027.42/30901. PMID 8458085. S2CID 802885. Archived from the original on 2020-03-13. Retrieved 2019-08-29.
- ^ Choi YB, Kadakkuzha BM, Liu XA, Akhmedov K, Kandel ER, Puthanveettil SV (July 23, 2014). "Huntingtin is critical both pre- and postsynaptically for long-term learning-related synaptic plasticity in Aplysia". PLOS ONE. 9 (7): e103004. Bibcode:2014PLoSO...9j3004C. doi:10.1371/journal.pone.0103004. PMC 4108396. PMID 25054562.
- ^ Nance MA, Mathias-Hagen V, Breningstall G, Wick MJ, McGlennen RC (Jan 1999). "Analysis of a very large trinucleotide repeat in a patient with juvenile Huntington's disease". Neurology. 52 (2): 392–4. doi:10.1212/wnl.52.2.392. PMID 9932964. S2CID 33091017. Archived from the original on 2009-05-05. Retrieved 2009-05-02.
- ^ Nasir J, Floresco SB, O'Kusky JR, Diewert VM, Richman JM, Zeisler J, Borowski A, Marth JD, Phillips AG, Hayden MR (Jun 1995). "Targeted disruption of the Huntington's disease gene results in embryonic lethality and behavioral and morphological changes in heterozygotes". Cell. 81 (5): 811–23. doi:10.1016/0092-8674(95)90542-1. PMID 7774020. S2CID 16835259.